II.3.1 Current understanding of the gene as a cistron is practical for the needs of molecular biology but is inadequate for the study of evolutionary processes.
Modern molecular biologists almost universally employ the term gene to denote a cistron. Cistron was originally defined in terms of the “cis-trans-test” (Fig. II-8). In this test, two mutants with mutations manifested in the same way in the phenotype of their carriers are first prepared or isolated from nature. If we are interested in whether both mutations are found in the same cistron, then a sufficiently long continuous DNA section bearing the mutation is transferred artificially, e.g. through transfection, or naturally, i.e. through crossing, from one mutant to cells containing the DNA of the individual with the other mutation. The obtained individual carries the given section in two copies, each of which bears one mutation. If the functioning of the particular gene is renewed in it, i.e. the original form of the trait corresponding to the unmutated form of the given gene is formed, it will be very probable that the relevant mutation will be located on two different cistrons. In the opposite case, this will correspond to a mutation on a single cistron. This test can be employed in this simple manner only if the wild form of the trait is dominant, i.e. when the presence of only a single copy of the given gene in the genome is required for its occurrence. Simultaneously, it is necessary to exclude (technically in advance or at least subsequently when evaluating the data) the
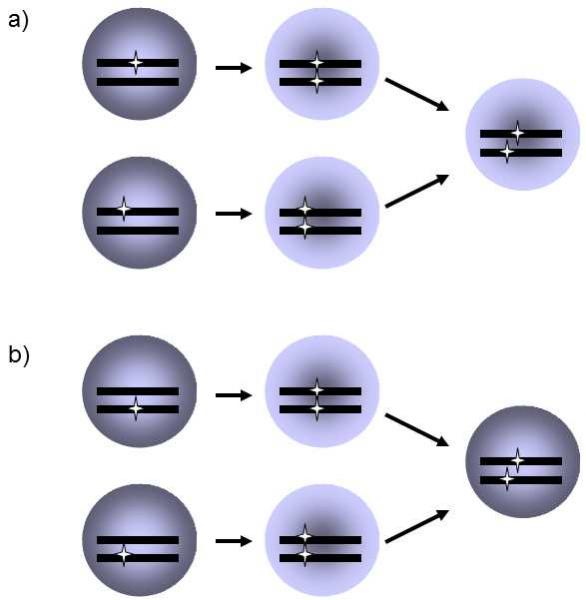
Fig. II.8 Cis-trans-test. Two recessive mutations that have the same phenotype manifestation can be located either in the same cistron or in two different cistrons. In the former case, the introduction of a section of a mutated allele derived from one of the mutants into the genome of the other mutant does not lead to elimination of the phenotype manifestation of the mutation (a); in contrast, it frequently leads to this in the second case (b). In experiments, it is necessary to employ mutants with a nonfunctional genetic recombination apparatus, as recombination in the section between the two mutations could lead to the formation of one molecule containing both mutations and one molecule without a mutation. In this case, the unmutated phenotype would probably be renewed.
possibility that the functioning of the given gene was renewed as a consequence of genetic recombination occurring directly in the studied DNA section. When a gene is identified with a cistron, a gene basically corresponds to a continuous DNA section coding, for example, a certain RNA chain, e.g. ribosomal RNA or mRNA coding a particular protein. The particular DNA section can be subsequently modified, e.g. at the level of mRNA, by cis-splicing, i.e. splicing and reconnection of its individual sections, but not by trans-splicing, i.e. connection of RNA sections derived from other RNA molecules that are rewritten from other DNA sections.
The concept of a gene as a cistron is very practical from the viewpoint of molecular biology. It permits more or less exact and particularly unambiguous delimitation of the genes coding the individual molecules that participate in the life processes of cells and multi-cellular organisms. As, at the present time, the study of these molecules forms the major content of the work of the greatest number of scientific workers in the field of biology, this conception of a gene quite predominates. It is seen by a great many biologists as quite obvious, correct and, in fact, the only possibility. However, this concept of a gene is quite inadequate for the purposes of evolutionary biology. It is apparent that two independent mutations at two places on a single cistron can have different, mutually independent phenotype manifestations. In sexually reproducing organisms, i.e. in most of currently known species, genetic recombination can occur at any time between these mutations in this section, which would physically separate not only the two mutations, but also the evolutionary fates of these mutations. Basically, every nucleotide in the DNA, to be more exact in the regulation and coding areas of the DNA, can thus act as an independent gene, can have a phenotype manifestation and can be transferred from one generation to the next. Whether two mutations in a single DNA chain will behave in evolution as two independent genes or as a single gene is decided by their mutual distance, or rather by the probability of their separation as a consequence of crossing-over, the probability that they will be passed down to the next generation, determined most frequently by the intensity of selection against their carriers or to their benefit, and also by the effective and nominal size of the population in which the evolution is occurring.
