VI.2.3 The effects of molecular drive are most obvious in the evolution of repetitive sequences in related species.
The existence of molecular drive is most clearly manifested in the evolution of repetitive DNA segments in closely related species (Charlesworth, Sniegowski, & Stephan 1994; Petes & Fink 1982). These segments are frequently located in the genome in a great many copies, of the order of hundreds of thousands. The individual copies are very similar and frequently completely identical. Simultaneously, repetitive sequences in closely related species are very different. It is difficult to explain this phenomenon without postulating the existence of a specific mechanism capable, following the speciation event – after the splitting off of a new species, of causing parallel differentiation in the repetitive DNA segments in all the loci of the genome. As the speciation process can hardly cause or affect the differentiation of repetitive genes, it is more reasonable to assume that this process occurs continuously in the gene pool of each species. Speciation division of the originally uniform gene pool into two gene pools alone only makes this visible, i.e. permits the repetitive sequences in the two gene pools to develop in different directions.
At the present time, it is mostly assumed that a random process of differentiation of repetitive genes occurs continuously in the gene pool of organisms and thus that mutations are accumulated in the individual copies of the repetitive gene. However, the process of homogenization of the individual copies also occurs simultaneously, i.e. a process in which the variants of the repetitive genes that are most successful from the standpoint of replication, transposition or gene conversion proliferate in the genome at the expense of other variants. In sexually reproducing organisms, the process of homogenization exceeds the boundaries of a single genome and the most successful variant of the repetitive sequence gradually proliferates in the whole gene pool. This is certainly a long-term process; however, it is relatively rapid compared to other evolutionary processes (Fig. VI.9). New variants of repetitive sequences become fixed in a substantially shorter time than the
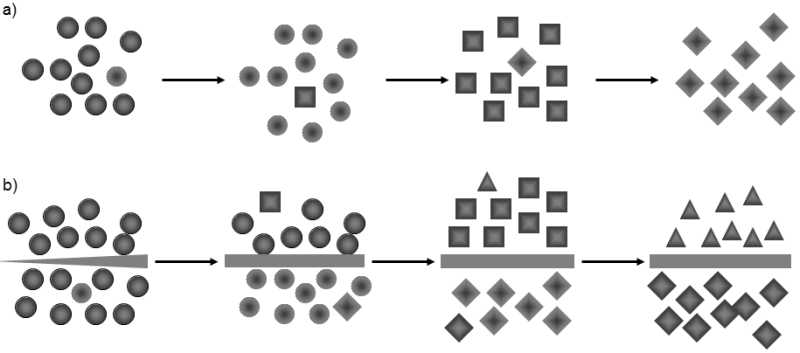
Fig. VI.9. Evolution of repetitive sequences in sister species. The evolution of repetitive sequences through the action of molecular drive occurs much more rapidly than speciation. Consequently, new variants of repetitive sequences are repeatedly fixed during the existence of a species. If the gene pool consists of a single unit (a), rapid fixation occurs in all the individuals of the given species, so that molecular-taxonomic studies mostly do not reveal the existence of polymorphism in repetitive sequences. However, as soon as speciation (b) occurs in the species, the original single gene pool divides into two parts and, from this moment, new variants of independent repetitive sequences are fixed in the two parts by molecular drive. Thus very rapid differentiation of the repetitive sequences occurs in both species.
interval separating two subsequent speciation events so that, when studying even closely related species, we find that different variants of the repetitive sequence became fixed in each of them.This fact can be utilized in molecular taxonomy – study of repetitive genes enables discrimination amongst representatives of even very closely related species (Grechko et al. 1998)(see also XXIV.3.9).
